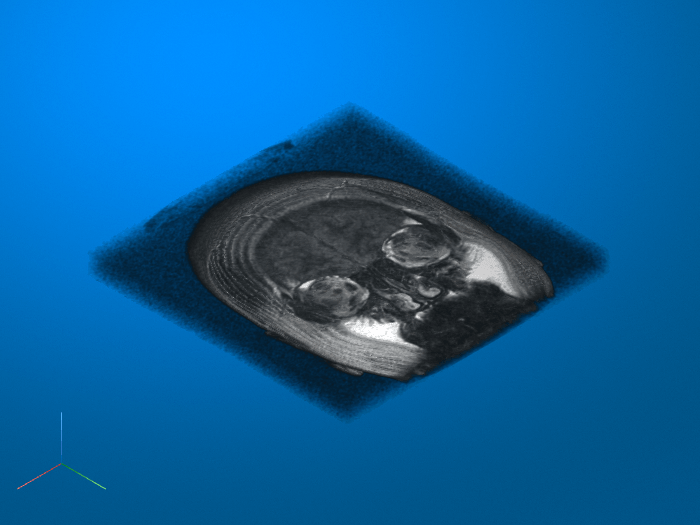
imsegkmeans3
K-means clustering based volume segmentation
Description
Examples
Input Arguments
Name-Value Arguments
Output Arguments
Tips
The function yields reproducible results. The output will not vary in multiple runs given the same input arguments.
References
[1]
Version History
Introduced in R2018b