addRelativePose
Add relative pose to pose graph
Syntax
Description
addRelativePose(
creates a node based on the input poseGraph,measurement)measurement that connects to
the last pose node in the pose graph. To add landmark nodes, see the
addPointLandmark function.
addRelativePose(
also specifies the information matrix as part of the edge constraint, which
represents the uncertainty of the pose measurement.poseGraph,measurement,infoMat)
addRelativePose(
creates a new pose node and connects it to the specific node specified by
poseGraph,measurement,infoMat,fromNodeID)fromNodeID.
addRelativePose(
creates an edge by specifying a relative pose measurement between existing nodes
specified by poseGraph,measurement,infoMat,fromNodeID,toNodeID)fromNodeID and toNodeID. This
edge is called a loop closure. If a loop closure already
exists, the function appends the new measurement. Calling the optimizePoseGraph function combines multiple appended measurements
into a single edge. This syntax does not support adding edges to a landmark
node.
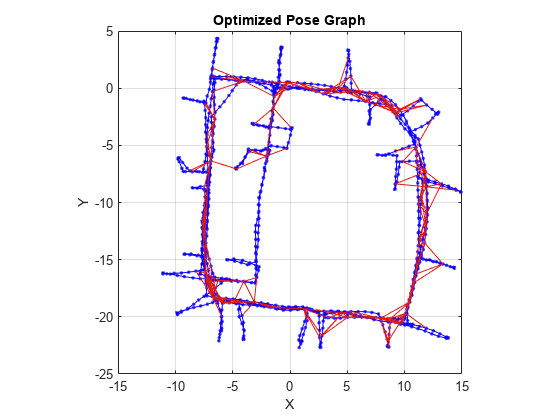
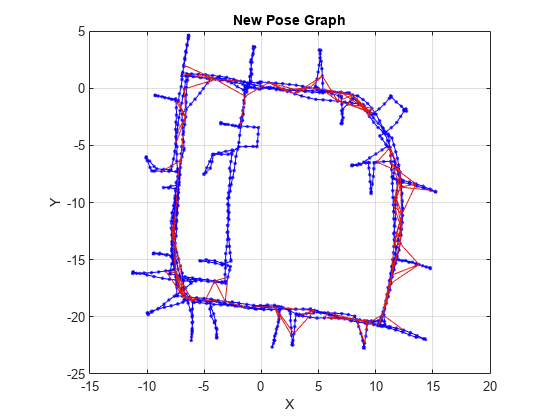
Examples
Input Arguments
Output Arguments
Extended Capabilities
Version History
Introduced in R2019b