Load the Target-Mediated Drug Disposition (TMDD) model.
Set the target occupancy (TO) as a response.
Get the dosing information.
Scan over different dose amounts using a SimBiology.Scenarios object. To do so, first parameterize the Amount property of the dose. Then vary the corresponding parameter value using the Scenarios object.
Create a SimFunction to simulate the model. Set TO as the simulation output.
f =
SimFunction
Parameters:
Name Value Type Units
_______________ _____ _____________ ____________
{'AmountParam'} 1 {'parameter'} {'nanomole'}
Observables:
Name Type Units
______ _____________ _________________
{'TO'} {'parameter'} {'dimensionless'}
Dosed:
TargetName TargetDimension Amount AmountValue AmountUnits
_______________ ___________________________________ _______________ ___________ ____________
{'Plasma.Drug'} {'Amount (e.g., mole or molecule)'} {'AmountParam'} 1 {'nanomole'}
TimeUnits: day
Simulate the model using the dose amounts generated by the Scenarios object. In this case, the object generates 31 different doses; hence the model is simulated 31 times and generates a SimData array.
SimBiology Simulation Data Array: 31-by-1
ModelName: TMDD
Logged Data:
Species: 0
Compartment: 0
Parameter: 1
Sensitivity: 0
Observable: 0
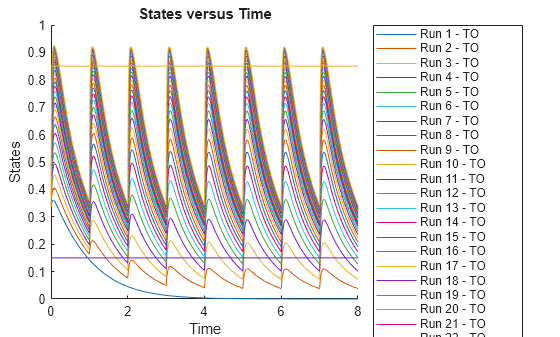
Plot the simulation results. Also add two reference lines that represent the safety and efficacy thresholds for TO. In this example, suppose that any TO value above 0.85 is unsafe, and any TO value below 0.15 has no efficacy.
Postprocess the simulation results. Find out which dose amounts are effective, corresponding to the TO responses within the safety and efficacy thresholds. To do so, add an observable expression to the simulation data.
SimBiology Simulation Data Array: 31-by-1
ModelName: TMDD
Logged Data:
Species: 0
Compartment: 0
Parameter: 1
Sensitivity: 0
Observable: 1
The addobservable function evaluates the new observable expression for each SimData in sd and returns the evaluated results as a new SimData array, newSD, which now has the added observable (stat1).
SimBiology stores the observable results in two different properties of a SimData object. If the results are scalar-valued, they are stored in SimData.ScalarObservables. Otherwise, they are stored in SimData.VectorObservables. In this example, the stat1 observable expression is scalar-valued.
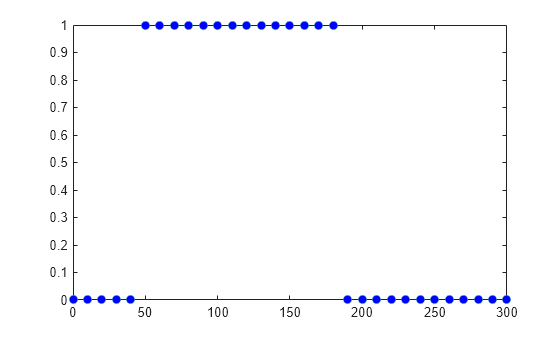
Extract the scalar observable values and plot them against the dose amounts.
The plot shows that dose amounts ranging from 50 to 180 nanomoles provide TO responses that lie within the target efficacy and safety thresholds.
You can update the observable expression with different threshold amounts. The function recalculates the expression and returns the results in a new SimData object array.
Rename the observable expression. The function renames the observable, updates any expressions that reference the renamed observable (if applicable), and returns the results in a new SimData object array.
Restore the warning settings.