refit
Refit neighborhood component analysis (NCA) model for regression
Description
mdlrefit = refit(mdl,Name=Value)mdl, with modified parameters specified by one
or more name-value arguments.
Examples
Load the sample data.
load("robotarm.mat")The robotarm (pumadyn32nm) data set is created using a robot arm simulator with 7168 training and 1024 test observations with 32 features [1], [2]. This is a preprocessed version of the original data set. Data are preprocessed by subtracting off a linear regression fit followed by normalization of all features to unit variance.
Compute the generalization error without feature selection.
nca = fsrnca(Xtrain,ytrain,FitMethod="none", ... Standardize=true); L = loss(nca,Xtest,ytest)
L = 0.9017
Now, refit the model and compute the prediction loss with feature selection, with = 0 (no regularization term) and compare to the previous loss value, to determine feature selection seems necessary for this problem. For the settings that you do not change, refit uses the settings of the initial model nca. For example, it uses the feature weights found in nca as the initial feature weights.
nca2 = refit(nca,FitMethod="exact",Lambda=0);
L2 = loss(nca2,Xtest,ytest)L2 = 0.1088
The decrease in the loss suggests that feature selection is necessary.
Plot the feature weights.
plot(nca2.FeatureWeights,"o")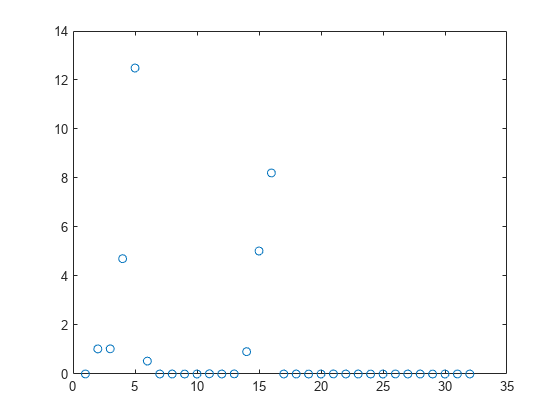
Tuning the regularization parameter usually improves the results. Suppose that, after tuning using cross-validation as in Tune Regularization Parameter in NCA for Regression, the best value found is 0.0035. Refit the nca model using this value and stochastic gradient descent as the solver. Compute the prediction loss.
nca3 = refit(nca2,FitMethod="exact",Lambda=0.0035, ... Solver="sgd"); L3 = loss(nca3,Xtest,ytest)
L3 = 0.0573
Plot the feature weights.
plot(nca3.FeatureWeights,"o")
After tuning the regularization parameter, the loss decreased even more and the software identified four of the features as relevant.
References
[1] Rasmussen, C. E., R. M. Neal, G. E. Hinton, D. van Camp, M. Revow, Z. Ghahramani, R. Kustra, and R. Tibshirani. The DELVE Manual, 1996, https://mlg.eng.cam.ac.uk/pub/pdf/RasNeaHinetal96.pdf
Input Arguments
Neighborhood component analysis model or classification, specified
as a FeatureSelectionNCARegression object.
Name-Value Arguments
Specify optional pairs of arguments as
Name1=Value1,...,NameN=ValueN, where Name is
the argument name and Value is the corresponding value.
Name-value arguments must appear after other arguments, but the order of the
pairs does not matter.
Example: refit(mdl,Lambda=0.01) refits the model
mdl with a lambda value of
0.01.
Fitting Options
Method for fitting the model, specified as one of the following.
"exact"— Performs fitting using all of the data."none"— No fitting. Use this option to evaluate the generalization error of the NCA model using the initial feature weights supplied in the call tofsrnca."average"— The function divides the data into partitions (subsets), fits each partition using theexactmethod, and returns the average of the feature weights. You can specify the number of partitions using theNumPartitionsname-value argument.
Example: FitMethod="none"
Regularization parameter, specified as a nonnegative scalar value.
For n observations, the best Lambda value
that minimizes the generalization error of the NCA model is expected
to be a multiple of 1/n
Example: Lambda=0.01
Data Types: double | single
Solver type for estimating feature weights, specified as one of the following.
"lbfgs"— Limited memory BFGS (Broyden-Fletcher-Goldfarb-Shanno) algorithm (LBFGS algorithm)"sgd"— Stochastic gradient descent"minibatch-lbfgs"— Stochastic gradient descent with LBFGS algorithm applied to mini-batches
Example: Solver="minibatch-lbfgs"
Initial feature weights, specified as a p-by-1 vector of real positive scalar values.
Data Types: double | single
Indicator for verbosity level for the convergence summary display, specified as one of the following.
0 — No convergence summary
1 — Convergence summary including iteration number, norm of the gradient, and objective function value.
>1 — More convergence information depending on the fitting algorithm
When using solver
"minibatch-lbfgs"and verbosity level >1, the convergence information includes iteration log from intermediate mini-batch LBFGS fits.
Example: Verbose=2
Data Types: double | single
LBFGS or Mini-Batch LBFGS Options
Relative convergence tolerance on the gradient norm for solver lbfgs,
specified as a positive real scalar value.
Example: GradientTolerance=0.00001
Data Types: double | single
SGD or Mini-Batch LBFGS Options
Initial learning rate for solver sgd, specified as a positive scalar
value.
When using solver type "sgd", the learning rate decays over iterations
starting with the value specified for InitialLearningRate.
Example: InitialLearningRate=0.8
Data Types: double | single
Maximum number of passes for solver "sgd" (stochastic gradient
descent), specified as a positive integer value. Every pass processes
size(mdl.X,1) observations.
Example: PassLimit=10
Data Types: double | single
SGD or LBFGS or Mini-Batch LBFGS Options
Maximum number of iterations, specified as a positive integer.
Example: IterationLimit=250
Data Types: double | single
Output Arguments
Neighborhood component analysis model or classification, returned as a FeatureSelectionNCARegression object. You can either save the
results as a new model or update the existing model as mdl =
refit(mdl,Name=Value).
Version History
Introduced in R2016b
See Also
FeatureSelectionNCARegression | loss | fsrnca | predict | selectFeatures
MATLAB Command
You clicked a link that corresponds to this MATLAB command:
Run the command by entering it in the MATLAB Command Window. Web browsers do not support MATLAB commands.
Select a Web Site
Choose a web site to get translated content where available and see local events and offers. Based on your location, we recommend that you select: .
You can also select a web site from the following list
How to Get Best Site Performance
Select the China site (in Chinese or English) for best site performance. Other MathWorks country sites are not optimized for visits from your location.
Americas
- América Latina (Español)
- Canada (English)
- United States (English)
Europe
- Belgium (English)
- Denmark (English)
- Deutschland (Deutsch)
- España (Español)
- Finland (English)
- France (Français)
- Ireland (English)
- Italia (Italiano)
- Luxembourg (English)
- Netherlands (English)
- Norway (English)
- Österreich (Deutsch)
- Portugal (English)
- Sweden (English)
- Switzerland
- United Kingdom (English)