loss
Regression error for Gaussian process regression model
Description
Examples
Load the sample data.
load('gprdata.mat')The data has 8 predictor variables and contains 500 observations in training data and 100 observations in test data. This is simulated data.
Fit a GPR model using the squared exponential kernel function with separate length scales for each predictor. Standardize the predictor values in the training data. Use the exact method for fitting and prediction.
gprMdl = fitrgp(Xtrain,ytrain,'FitMethod','exact',... 'PredictMethod','exact','KernelFunction','ardsquaredexponential',... 'Standardize',1);
Compute the regression error for the test data.
L = loss(gprMdl,Xtest,ytest)
L = 0.6928
Predict the responses for test data.
ypredtest = predict(gprMdl,Xtest);
Plot the test response along with the predictions.
figure; plot(ytest,'r'); hold on; plot(ypredtest,'b'); legend('Data','Predictions','Location','Best');
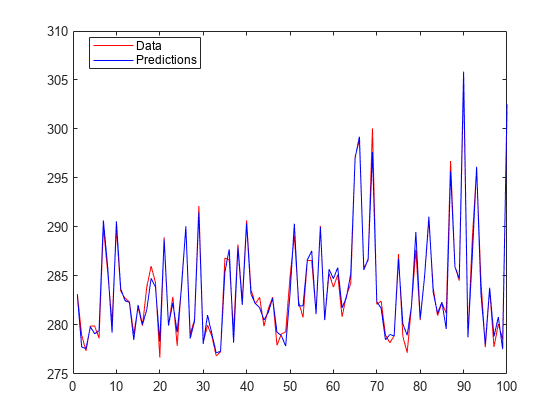
Manually compute the regression loss.
L = (ytest - ypredtest)'*(ytest - ypredtest)/length(ytest)
L = 0.6928
Load the sample data and store in a table.
load fisheriris tbl = table(meas(:,1),meas(:,2),meas(:,3),meas(:,4),species,... 'VariableNames',{'meas1','meas2','meas3','meas4','species'});
Fit a GPR model using the first measurement as the response and the other variables as the predictors.
mdl = fitrgp(tbl,'meas1');Predict the responses using the trained model.
ypred = predict(mdl,tbl);
Compute the mean absolute error.
n = height(tbl);
y = tbl.meas1;
fun = @(y,ypred,w) sum(abs(y-ypred))/n;
L = loss(mdl,tbl,'lossfun',fun)L = 0.2345
Input Arguments
Gaussian process regression model, specified as a RegressionGP
(full) or CompactRegressionGP (compact) object.
New data, specified as a table or an
n-by-d matrix, where m is the
number of observations, and d is the number of predictor variables in
the training data.
If you trained gprMdl on a table, then
Xnew must be a table that contains all the
predictor variables used to train gprMdl.
If Xnew is a table, then it can also contain
Ynew. And if it does, then you do not have to specify
Ynew.
If you trained gprMdl on a matrix, then
Xnew must be a numeric matrix with d columns,
and can only contain values for the predictor variables.
Data Types: single | double | table
New observed response values, that correspond to the predictor values in
Xnew, specified as an n-by-1 vector.
n is the number of rows in Xnew. Each entry in
Ynew is the observed response based on the predictor data in the
corresponding row of Xnew.
If Xnew is a table containing new response values, you do not
have to specify Ynew.
Data Types: single | double
Name-Value Arguments
Specify optional pairs of arguments as
Name1=Value1,...,NameN=ValueN, where Name is
the argument name and Value is the corresponding value.
Name-value arguments must appear after other arguments, but the order of the
pairs does not matter.
Before R2021a, use commas to separate each name and value, and enclose
Name in quotes.
Example: loss(grpMdl,Xnew,Ynew,"PredictionForMissingValue","omitted")
specifies to omit observations with missing prediction values from the loss
computation.
Loss function, specified as 'mse' (mean squared error) or a
function handle.
If you pass a function handle, say fun, loss calls it as shown below: fun(Y,Ypred,W), where
Y, Ypred and W are
numeric vectors of length n, and n is the number
of rows in Xnew. Y is the observed response,
Ypred is the predicted response, and W is
the observation weights.
Example: 'lossfun',Fct calls the loss function
Fct.
Data Types: char | string | function_handle
Since R2023b
Predicted response value to use for observations with missing predictor values,
specified as "median", "mean",
"omitted", or a numeric scalar.
| Value | Description |
|---|---|
"median" | loss uses the median of the observed
response values in the training data as the predicted response value for
observations with missing predictor values. |
"mean" | loss uses the mean of the observed
response values in the training data as the predicted response value for
observations with missing predictor values. |
"omitted" | loss excludes observations with missing
predictor values from the loss computation. |
| Numeric scalar | loss uses this value as the predicted
response value for observations with missing predictor values. |
If an observation is missing an observed response value or an observation weight, then
loss does not use the observation in the loss
computation.
Example: PredictionForMissingValue="omitted"
Data Types: single | double | char | string
Observation weights, specified as n-by-1 vector, where
n is the number of rows in Xnew. By default,
the weight of each observation is 1.
Example: 'weights',W uses the observation weights in vector
W.
Data Types: double | single
Output Arguments
Regression error for the trained Gaussian process regression model,
gprMdl, returned as a scalar value.
Alternatives
You can use resubLoss to
compute the regression error for the trained GPR model at the observations in the training
data.
Extended Capabilities
The
loss function fully supports tall arrays. For more information,
see Tall Arrays.
This function fully supports GPU arrays. For more information, see Run MATLAB Functions on a GPU (Parallel Computing Toolbox).
Version History
Introduced in R2015bloss fully supports GPU arrays.
Starting in R2023b, when you predict or compute the loss, some regression models allow you to specify the predicted response value for observations with missing predictor values. Specify the PredictionForMissingValue name-value argument to use a numeric scalar, the training set median, or the training set mean as the predicted value. When computing the loss, you can also specify to omit observations with missing predictor values.
This table lists the object functions that support the
PredictionForMissingValue name-value argument. By default, the
functions use the training set median as the predicted response value for observations with
missing predictor values.
| Model Type | Model Objects | Object Functions |
|---|---|---|
| Gaussian process regression (GPR) model | RegressionGP, CompactRegressionGP | loss, predict, resubLoss, resubPredict |
RegressionPartitionedGP | kfoldLoss, kfoldPredict | |
| Gaussian kernel regression model | RegressionKernel | loss, predict |
RegressionPartitionedKernel | kfoldLoss, kfoldPredict | |
| Linear regression model | RegressionLinear | loss, predict |
RegressionPartitionedLinear | kfoldLoss, kfoldPredict | |
| Neural network regression model | RegressionNeuralNetwork, CompactRegressionNeuralNetwork | loss, predict, resubLoss, resubPredict |
RegressionPartitionedNeuralNetwork | kfoldLoss, kfoldPredict | |
| Support vector machine (SVM) regression model | RegressionSVM, CompactRegressionSVM | loss, predict, resubLoss, resubPredict |
RegressionPartitionedSVM | kfoldLoss, kfoldPredict |
In previous releases, the regression model loss and predict functions listed above used NaN predicted response values for observations with missing predictor values. The software omitted observations with missing predictor values from the resubstitution ("resub") and cross-validation ("kfold") computations for prediction and loss.
The loss function no longer omits an observation with a
NaN prediction when computing the weighted average regression loss. Therefore,
loss can now return NaN when the predictor data
Xnew contains any missing values. In most cases, if the test set
observations do not contain missing predictors, the loss
function does not return NaN.
This change improves the automatic selection of a regression model when you use
fitrauto.
Before this change, the software might select a model (expected to best predict the
responses for new data) with few non-NaN predictors.
If loss in your code returns NaN, you can update your code
to avoid this result. Remove or replace the missing values by using rmmissing or fillmissing, respectively.
The following table shows the regression models for which the
loss object function might return NaN. For more details, see
the Compatibility Considerations for each loss
function.
| Model Type | Full or Compact Model Object | loss Object Function |
|---|---|---|
| Gaussian process regression (GPR) model | RegressionGP, CompactRegressionGP | loss |
| Gaussian kernel regression model | RegressionKernel | loss |
| Linear regression model | RegressionLinear | loss |
| Neural network regression model | RegressionNeuralNetwork, CompactRegressionNeuralNetwork | loss |
| Support vector machine (SVM) regression model | RegressionSVM, CompactRegressionSVM | loss |
See Also
fitrgp | RegressionGP | CompactRegressionGP | compact | resubLoss | predict
MATLAB Command
You clicked a link that corresponds to this MATLAB command:
Run the command by entering it in the MATLAB Command Window. Web browsers do not support MATLAB commands.
Select a Web Site
Choose a web site to get translated content where available and see local events and offers. Based on your location, we recommend that you select: .
You can also select a web site from the following list
How to Get Best Site Performance
Select the China site (in Chinese or English) for best site performance. Other MathWorks country sites are not optimized for visits from your location.
Americas
- América Latina (Español)
- Canada (English)
- United States (English)
Europe
- Belgium (English)
- Denmark (English)
- Deutschland (Deutsch)
- España (Español)
- Finland (English)
- France (Français)
- Ireland (English)
- Italia (Italiano)
- Luxembourg (English)
- Netherlands (English)
- Norway (English)
- Österreich (Deutsch)
- Portugal (English)
- Sweden (English)
- Switzerland
- United Kingdom (English)