infer
Infer conditional variances of conditional variance models
Description
Tbl2 = infer(Mdl,Tbl1)Tbl2 containing the inferred
conditional variances and innovations from evaluating the fully specified,
univariate conditional variance model Mdl at the response
variable data in the table or timetable Tbl1. When
Mdl is a model fitted to the response data and returned by
estimate, the inferred innovations are
residuals. (since R2023a)
infer selects the response variable named in
Mdl.SeriesName or the sole variable in
Tbl1. To select a different response variable in
Tbl1 at which to evaluate the model, use the
ResponseVariable name-value argument.
___
= infer(___,
specifies options using one or more name-value arguments in
addition to any of the input argument combinations in previous syntaxes.
Name,Value)infer returns the output argument combination for the
corresponding input arguments. For example, infer(Mdl,Y,V0=v0) initializes the
conditional variance model of Mdl using the presample
conditional variance data in v0.
Examples
Infer conditional variances from a GARCH(1,1) model with known coefficients. Specify response data as a numeric vector.
Specify a GARCH(1,1) model with known parameters. Simulate 101 conditional variances and responses (innovations) from the model. Set aside the first observation from each series to use as presample data.
Mdl = garch(Constant=0.01,GARCH=0.8,ARCH=0.15); rng(1,"twister") % For reproducibility [vS,yS] = simulate(Mdl,101); y0 = yS(1); v0 = vS(1); y = yS(2:end); v = vS(2:end); figure tiledlayout(2,1) nexttile plot(v) title("Conditional Variances") nexttile plot(y) title("Innovations")

Infer the conditional variances of y without using presample data. Compare them to the known (simulated) conditional variances.
vI = infer(Mdl,y); figure plot(1:100,v,"r",LineWidth=2) hold on plot(1:100,vI,"k:",LineWidth=1.5) legend("Simulated","Inferred",Location="northeast") title("Inferred Conditional Variances, No Presample") hold off

Notice the transient response (discrepancy) in the early time periods due to the absence of presample data.
Infer conditional variances using the set-aside presample innovation, y0. Compare them to the known (simulated) conditional variances.
vE = infer(Mdl,y,E0=y0); figure plot(1:100,v,"r",LineWidth=2) hold on plot(1:100,vE,"k:",LineWidth=1.5) legend("Simulated","Inferred",Location="northeast") title("Inferred Conditional Variances, Innovations Presample") hold off

There is a slightly reduced transient response in the early time periods.
Infer conditional variances using the presample of conditional variance data, v0. Compare them to the known (simulated) conditional variances.
vO = infer(Mdl,y,V0=v0); figure plot(v) hold on plot(1:100,v,"r",LineWidth=2) plot(1:100,vO,"k:",LineWidth=1.5) legend("Simulated","Inferred",Location="northeast") title("Inferred Conditional Variances, Conditional Variance Presample") hold off

There is a much smaller transient response in the early time periods.
Infer conditional variances using both the presample innovation and conditional variance. Compare them to the known (simulated) conditional variances.
vEO = infer(Mdl,y,E0=y0,V0=v0); figure plot(v) hold on plot(1:100,v,"r",LineWidth=2) plot(1:100,vEO,"k:",LineWidth=1.5) legend("Simulated","Inferred",Location="northeast") title("Inferred Conditional Variances, Presamples") hold off

When you use sufficient presample innovations and conditional variances, the inferred conditional variances are exact (there is no transient response).
Infer conditional variances from an EGARCH(1,1) model with known coefficients. When you use, and then do not use presample data, compare the results from infer.
Specify an EGARCH(1,1) model with known parameters. Simulate 101 conditional variances and responses (innovations) from the model. Set aside the first observation from each series to use as presample data.
Mdl = egarch(Constant=0.001,GARCH=0.8, ... ARCH=0.15,Leverage=-0.1); rng(1,"twister") % For reproducibility [vS,yS] = simulate(Mdl,101); y0 = yS(1); v0 = vS(1); y = yS(2:end); v = vS(2:end); figure tiledlayout(2,1) nexttile plot(v) title("Conditional Variances") nexttile plot(y) title("Innovations")

Infer the conditional variances of y without using any presample data. Compare them to the known (simulated) conditional variances.
vI = infer(Mdl,y); figure plot(1:100,v,"r",LineWidth=2) hold on plot(1:100,vI,"k:",LineWidth=1.5) legend("Simulated","Inferred",Location="northeast") title("Inferred Conditional Variances, No Presample") hold off
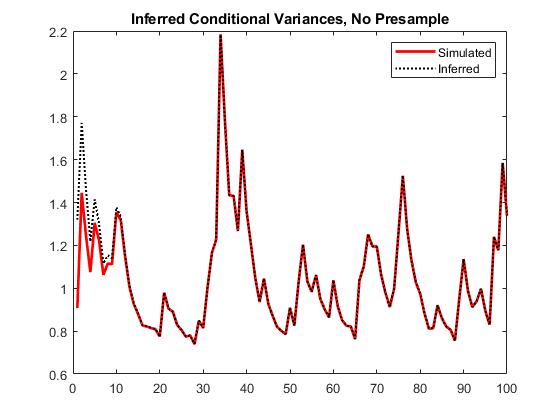
Notice the transient response (discrepancy) in the early time periods due to the absence of presample data.
Infer conditional variances using the set-aside presample innovation, y0. Compare them to the known (simulated) conditional variances.
vE = infer(Mdl,y,E0=y0); figure plot(1:100,v,"r",LineWidth=2) hold on plot(1:100,vE,"k:",LineWidth=1.5) legend("Simulated","Inferred",Location="northeast") title("Inferred Conditional Variances, Presample E") hold off

There is a slightly reduced transient response in the early time periods.
Infer conditional variances using the set-aside presample variance, v0. Compare them to the known (simulated) conditional variances.
vO = infer(Mdl,y,V0=v0); figure plot(v) hold on plot(1:100,v,"r",LineWidth=2) plot(1:100,vO,"k:",LineWidth=1.5) legend("Simulated","Inferred",Location="northeast") title("Inferred Conditional Variances, Presample V") hold off
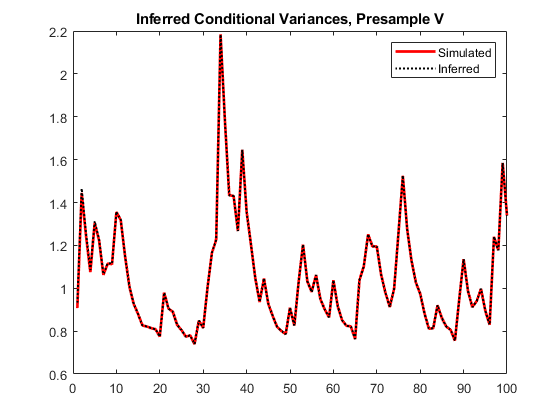
The transient response is almost eliminated.
Infer conditional variances using both the presample innovation and conditional variance. Compare them to the known (simulated) conditional variances.
vEO = infer(Mdl,y,E0=y0,V0=v0); figure plot(v) hold on plot(1:100,v,"r",LineWidth=2) plot(1:100,vEO,"k:",LineWidth=1.5) legend("Simulated","Inferred",Location="northeast") title("Inferred Conditional Variances, Presamples") hold off
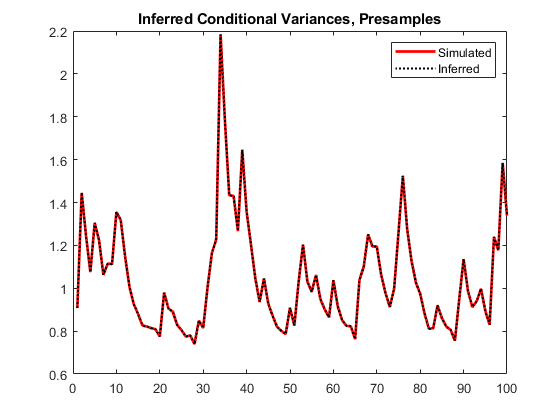
When you use sufficient presample innovations and conditional variances, the inferred conditional variances are exact (there is no transient response).
Infer conditional variances from a GJR(1,1) model with known coefficients. When you use, and then do not use presample data, compare the results from infer.
Specify a GJR(1,1) model with known parameters. Simulate 101 conditional variances and responses (innovations) from the model. Set aside the first observation from each series to use as presample data.
Mdl = gjr(Constant=0.01,GARCH=0.8,ARCH=0.14,Leverage=0.1); rng(1,"twister") % For reproducibility [vS,yS] = simulate(Mdl,101); y0 = yS(1); v0 = vS(1); y = yS(2:end); v = vS(2:end); figure tiledlayout(2,1) nexttile plot(v) title("Conditional Variances") nexttile plot(y) title("Innovations")
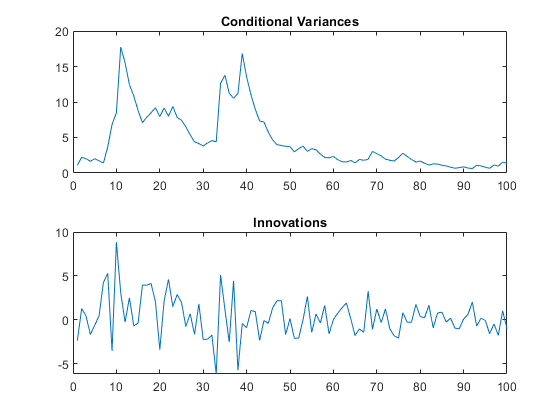
Infer the conditional variances of y without using any presample data. Compare them to the known (simulated) conditional variances.
vI = infer(Mdl,y); figure plot(1:100,v,LineWidth=2) hold on plot(1:100,vI,":",LineWidth=1.5) legend("Simulated","Inferred",Location="northeast") title("Inferred Conditional Variances, No Presample") hold off
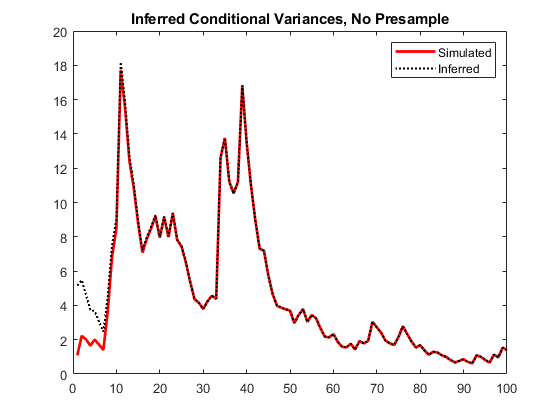
Notice the transient response (discrepancy) in the early time periods due to the absence of presample data.
Infer conditional variances using the set-aside presample innovation, y0. Compare them to the known (simulated) conditional variances.
vE = infer(Mdl,y,E0=y0); figure plot(1:100,v,LineWidth=2) hold on plot(1:100,vE,":",LineWidth=1.5) legend("Simulated","Inferred",Location="northeast") title("Inferred Conditional Variances, Presample E") hold off
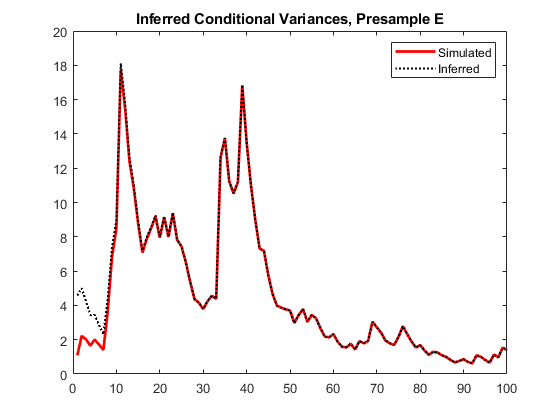
The provided presample innovations slightly reduces the transient response in the early time periods.
Infer conditional variances using the set-aside presample conditional variance, vO. Compare them to the known (simulated) conditional variances.
vO = infer(Mdl,y,V0=v0); figure plot(1:100,v,LineWidth=2) hold on plot(1:100,vO,":",LineWidth=1.5) legend("Simulated","Inferred",Location="northeast") title("Inferred Conditional Variances, Presample V") hold off
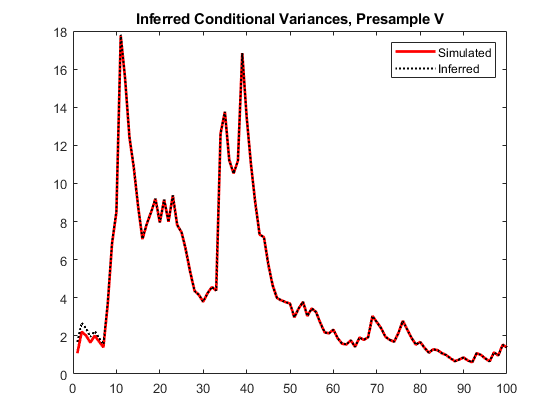
The provided presample conditional variances reduces the transient response in the early time periods more than the reduction resulting from providing presample innovations.
Infer conditional variances using both the presample innovation and conditional variance. Compare them to the known (simulated) conditional variances.
vEO = infer(Mdl,y,E0=y0,V0=v0); figure plot(1:100,v,LineWidth=2) hold on plot(1:100,vEO,":",LineWidth=1.5) legend("Simulated","Inferred",Location="northeast") title("Inferred Conditional Variances, Presamples") hold off

The provided presample innovations and conditional variances reduces the transient response in the early time periods to zero.
Since R2023a
Infer the loglikelihood objective function values for an EGARCH(1,1) and EGARCH(2,1) model fit to the average weekly closing NASDAQ returns. To identify which model is the more parsimonious, adequate fit, conduct a likelihood ratio test. Specify data in timetables.
Load the U.S. equity indices data Data_EquityIdx.mat.
load Data_EquityIdxThe timetable DataTimeTable contains the daily NASDAQ closing prices, among other indices.
Compute the weekly average closing prices of all timetable variables.
DTTW = convert2weekly(DataTimeTable,Aggregation="mean");Compute the weekly returns and their sample mean.
DTTRet = price2ret(DTTW); DTTRet.Interval = []; T = height(DTTRet)
T = 626
When you plan to supply a timetable, you must ensure it has all the following characteristics:
The selected response variable is numeric and does not contain any missing values.
The timestamps in the
Timevariable are regular, and they are ascending or descending.
Remove all missing values from the timetable, relative to the NASDAQ returns series.
DTTRet = rmmissing(DTTRet,DataVariables="NASDAQ");
numobs = height(DTTRet)numobs = 626
Because all sample times have observed NASDAQ returns, rmmissing does not remove any observations.
Determine whether the sampling timestamps have a regular frequency and are sorted.
areTimestampsRegular = isregular(DTTRet,"weeks")areTimestampsRegular = logical
1
areTimestampsSorted = issorted(DTTRet.Time)
areTimestampsSorted = logical
1
areTimestampsRegular = 1 indicates that the timestamps of DTTRet represent a regular weekly sample. areTimestampsSorted = 1 indicates that the timestamps are sorted.
Reserve the first two observations to use as a presample.
DTTRet0 = DTTRet(1:2,:); DTTRet = DTTRet(3:end,:);
Fit an EGARCH(1,1) model to the returns. Supply in-sample and presample data in timetables, and specify NASDAQ as the variable containing the presample innovations. Infer the loglikelihood objective function value.
MdlEGARCH11 = egarch(1,1); MdlEGARCH11.SeriesName = "NASDAQ"; EstMdlEGARCH11 = estimate(MdlEGARCH11,DTTRet, ... Presample=DTTRet0,PresampleInnovationVariable="NASDAQ");
EGARCH(1,1) Conditional Variance Model (Gaussian Distribution):
Value StandardError TStatistic PValue
________ _____________ __________ __________
Constant -0.48899 0.15218 -3.2133 0.0013123
GARCH{1} 0.95567 0.013348 71.598 0
ARCH{1} 0.2766 0.052276 5.2912 1.2154e-07
Leverage{1} -0.10593 0.025607 -4.1366 3.5244e-05
[TblEGARCH11,logLEGARCH11] = infer(EstMdlEGARCH11,DTTRet, ... Presample=DTTRet0,PresampleInnovationVariable="NASDAQ"); tail(TblEGARCH11)
Time NYSE NASDAQ NASDAQ_Variance NASDAQ_Residual
___________ ___________ ___________ _______________ _______________
16-Nov-2001 0.0021092 0.0048052 3.4439e-05 0.0048052
23-Nov-2001 0.001451 0.00085891 3.0717e-05 0.00085891
30-Nov-2001 -0.00039051 0.0020552 2.4587e-05 0.0020552
07-Dec-2001 0.00087108 0.005263 2.0775e-05 0.005263
14-Dec-2001 -0.002694 -0.0012244 2.0067e-05 -0.0012244
21-Dec-2001 0.0019929 -0.00094985 1.7698e-05 -0.00094985
28-Dec-2001 0.0019952 -4.93e-05 1.5413e-05 -4.93e-05
04-Jan-2002 -0.00011742 -0.0012263 1.2447e-05 -0.0012263
TblEGARCH11 is a timetable of NASDAQ residuals NASDAQ_Residual and conditional variances NASDAQ_Variance, and all variables in the specified in-sample data DTTRet. logLEGARCH11 is the loglikelihood of the estimated model EstMdlEGARCH11 evaluated at the specified presample and in-sample data.
Fit an EGARCH(2,1) model to the returns. Supply in-sample and presample data in timetables, and specify NASDAQ as the variable containing the presample innovations. Infer the loglikelihood objective function value.
MdlEGARCH21 = egarch(2,1); MdlEGARCH21.SeriesName = "NASDAQ"; EstMdlEGARCH21 = estimate(MdlEGARCH21,DTTRet, ... Presample=DTTRet0,PresampleInnovationVariable="NASDAQ");
EGARCH(2,1) Conditional Variance Model (Gaussian Distribution):
Value StandardError TStatistic PValue
_________ _____________ __________ __________
Constant -0.48624 0.15764 -3.0846 0.0020386
GARCH{1} 0.96844 0.27069 3.5777 0.00034661
GARCH{2} -0.012511 0.26614 -0.047009 0.96251
ARCH{1} 0.27421 0.074046 3.7032 0.00021289
Leverage{1} -0.10486 0.035239 -2.9756 0.0029237
[TblEGARCH21,logLEGARCH21] = infer(EstMdlEGARCH21,DTTRet, ... Presample=DTTRet0,PresampleInnovationVariable="NASDAQ");
Conduct a likelihood ratio test, with the more parsimonious EGARCH(1,1) model as the null model, and the EGARCH(2,1) model as the alternative. The degree of freedom for the test is 1, because the EGARCH(2,1) model has one more parameter than the EGARCH(1,1) model (an additional GARCH term).
[h,p] = lratiotest(logLEGARCH21,logLEGARCH11,1)
h = logical
0
p = 0.9566
The null hypothesis is not rejected (h = 0). At the 0.05 significance level, the EGARCH(1,1) model is not rejected in favor of the EGARCH(2,1) model.
A GARCH(P, Q) model is nested within a GJR(P, Q) model. Therefore, you can perform a likelihood ratio test to compare GARCH(P, Q) and GJR(P, Q) model fits.
Infer the loglikelihood objective function values for a GARCH(1,1) and GJR(1,1) model fit to NASDAQ Composite Index returns. Conduct a likelihood ratio test to identify which model is the more parsimonious, adequate fit.
Load the NASDAQ data included with the toolbox, and convert the index to returns. Set aside the first two observations to use as presample data.
load Data_EquityIdx
nasdaq = DataTable.NASDAQ;
r = price2ret(nasdaq);
r0 = r(1:2);
rn = r(3:end);Fit a GARCH(1,1) model to the returns, and infer the loglikelihood objective function value.
Mdl1 = garch(1,1); EstMdl1 = estimate(Mdl1,rn,E0=r0);
GARCH(1,1) Conditional Variance Model (Gaussian Distribution):
Value StandardError TStatistic PValue
__________ _____________ __________ __________
Constant 2.0348e-06 5.4545e-07 3.7305 0.00019111
GARCH{1} 0.88275 0.008499 103.87 0
ARCH{1} 0.10956 0.0076979 14.233 5.7259e-46
[~,logL1] = infer(EstMdl1,rn,E0=r0);
Fit a GJR(1,1) model to the returns, and infer the loglikelihood objective function value.
Mdl2 = gjr(1,1); EstMdl2 = estimate(Mdl2,rn,E0=r0);
GJR(1,1) Conditional Variance Model (Gaussian Distribution):
Value StandardError TStatistic PValue
__________ _____________ __________ __________
Constant 4.4353e-06 7.1313e-07 6.2194 4.9908e-10
GARCH{1} 0.80675 0.013843 58.279 0
ARCH{1} 0.11245 0.013469 8.3488 6.8985e-17
Leverage{1} 0.11493 0.014356 8.0053 1.1914e-15
[~,logL2] = infer(EstMdl2,rn,E0=r0);
Conduct a likelihood ratio test, with the more parsimonious GARCH(1,1) model as the null model, and the GJR(1,1) model as the alternative. The degree of freedom for the test is 1, because the GJR(1,1) model has one more parameter than the GARCH(1,1) model (a leverage term).
[h,p] = lratiotest(logL2,logL1,1)
h = logical
1
p = 1.2283e-05
The null hypothesis is rejected (h = 1). At the 0.05 significance level, the GARCH(1,1) model is rejected in favor of the GJR(1,1) model.
Input Arguments
Response data, specified as a numobs-by-1 numeric
column vector or numobs-by-numpaths
matrix.
As a column vector, Y represents a single path of the
underlying series.
As a matrix, the rows of Y correspond to periods and
the columns correspond to separate paths. The observations across any row
occur simultaneously.
infer infers the conditional variances of
Y. Y usually represents an
innovation series with mean 0 and variances characterized by
Mdl. It is the continuation of the presample
innovation series E0. Y can also
represent a time series of innovations with mean 0 plus an offset. If
Mdl has a nonzero offset, then the software stores
its value in the Offset property
(Mdl.Offset).
The last observation of any series is the latest observation.
Since R2023a
Time series data containing response variable
yt, at which
infer evaluates the conditional variance
model Mdl, specified as a table or timetable with
numvars variables and numobs rows.
You can optionally select a response variable by using the
ResponseVariable name-value argument.
The selected variable is a single path (numobs-by-1
vector) or multiple paths
(numobs-by-numpaths matrix) of
numobs observations of response data. Each row is an
observation, and measurements in each row occur simultaneously.
The selected response variable in Tbl1 is a
numobs-by-numpaths numeric matrix.
Each row is an observation, and measurements in each row occur
simultaneously.
Each path (column) of the selected variable is independent of the other paths.
If Tbl1 is a timetable, it must represent a sample
with a regular datetime time step (see isregular), and the datetime
vector Tbl1.Time must be strictly ascending or
descending.
If Tbl1 is a table, the last row contains the latest
observation.
Name-Value Arguments
Specify optional pairs of arguments as
Name1=Value1,...,NameN=ValueN, where Name is
the argument name and Value is the corresponding value.
Name-value arguments must appear after other arguments, but the order of the
pairs does not matter.
Before R2021a, use commas to separate each name and value, and enclose
Name in quotes.
Example: 'E0',[1 1;0.5 0.5],'V0',[1 0.5;1 0.5] specifies two
equivalent presample paths of innovations and two, different presample paths of
conditional variances.
Since R2023a
Variable to select from Tbl1 to treat as the response variable
yt, specified as one of the following
data types:
String scalar or character vector containing a variable name in
Tbl1.Properties.VariableNamesVariable index (integer) to select from
Tbl1.Properties.VariableNamesA length
numvarslogical vector, whereResponseVariable(selects variablej) = truejTbl1.Properties.VariableNames, andsum(ResponseVariable)is1
The selected variable must be a numeric vector and cannot contain missing values (NaN).
If Tbl1 has one variable, the default specifies that variable. Otherwise, the default matches the variable to name in Mdl.SeriesName.
Example: ResponseVariable="StockRate2"
Example: ResponseVariable=[false false true false] or ResponseVariable=3 selects the third table variable as the response variable.
Data Types: double | logical | char | cell | string
Presample innovation paths
εt, specified as a
numpreobs-by-1 numeric column vector or a
numpreobs-by-numprepaths
matrix. The presample innovations provide initial values for the
innovations process of the conditional variance model
Mdl, and derive from a distribution with mean
0. Use E0 only when you supply the numeric array of
response data Y.
numpreobs is the number of presample observations.
numprepaths is the number of presample response
paths.
Each row is a presample observation, and measurements in each row
occur simultaneously. The last row contains the latest presample
observation. numpreobs must be at least
Mdl.Q. If numpreobs >
Mdl.Q, infer uses the
latest required number of observations only. The last element or row
contains the latest observation.
If
E0is a column vector, it represents a single path of the underlying innovation series.inferapplies it to each output path.If
E0is a matrix, each column represents a presample path of the underlying innovation series.numprepathsmust be at leastnumpaths. Ifnumprepaths>numpaths,inferuses the firstsize(Y,2)columns only.
The defaults are:
For GARCH(P,Q) and GJR(P,Q) models,
infersets any necessary presample innovations to the square root of the average squared value of the offset-adjusted response seriesY.For EGARCH(P,Q) models,
infersets any necessary presample innovations to zero.
Data Types: double
Positive presample conditional variance paths
σt2,
specified as a numpreobs-by-1 positive column vector
or numpreobs-by-numprepaths
positive matrix. V0 provides initial values for the
conditional variances in the model. Use V0 only when
you supply the numeric array of disturbances
Z.
Each row is a presample observation, and measurements in each row occur simultaneously. The last row contains the latest presample observation.
For GARCH(P,Q) and GJR(P,Q) models,
numpreobsmust be at leastMdl.P.For EGARCH(P,Q) models,
numpreobsmust be at leastmax([Mdl.P Mdl.Q]).
numpreobs must be at least
max([Mdl.P Mdl.Q]). If
numpreobs > max([Mdl.P
Mdl.Q]), infer uses the latest
required number of observations only. The last element or row contains
the latest observation.
If
V0is a column vector, it represents a single path of the conditional variance series.inferapplies it to each output path.If
V0is a matrix, each column represents a presample path of the conditional variance series.numprepathsmust be at leastnumpaths. Ifnumprepaths>numpaths,inferuses the firstsize(Y,2)columns only.
By default, infer sets any necessary
presample conditional variances to the unconditional variance of the
process.
Data Types: double
Since R2023a
Presample data containing paths of innovation
εt or conditional
variance
σt2
series to initialize the model, specified as a table or timetable, the
same type as Tbl1, with
numprevars variables and
numpreobs rows. Use
Presample only when you supply a table or
timetable of data Tbl1.
Each selected variable is a single path
(numpreobs-by-1 vector) or multiple paths
(numpreobs-by-numprepaths
matrix) of numpreobs observations representing the
presample of numpreobs observations of the innovation
or conditional variance series for
ResponseVariable, the selected response variable in
Tbl1.
Each row is a presample observation, and measurements in each row
occur simultaneously. numpreobs must be one of the
following values:
Mdl.QwhenPresampleprovides only presample innovations.Mdl.PwhenPresampleprovides only presample conditional variances.max([Mdl.P Mdl.Q])whenPresampleprovides both presample innovations and conditional variances
If numpreobs exceeds the minimum
number, infer uses the latest required number
of observations only.
If Presample is a timetable, all the following
conditions must be true:
Presamplemust represent a sample with a regular datetime time step (seeisregular).The inputs
Tbl1andPresamplemust be consistent in time such thatPresampleimmediately precedesTbl1with respect to the sampling frequency and order.The datetime vector of sample timestamps
Presample.Timemust be ascending or descending.
If Presample is a table, the last row contains
the latest presample observation.
The defaults are:
For GARCH(P,Q) and GJR(P,Q) models,
infersets any necessary presample innovations to the square root of the average squared value of the offset-adjusted response seriesY.For EGARCH(P,Q) models,
infersets any necessary presample innovations to zero.infersets any necessary presample conditional variances to the unconditional variance of the process.
If you specify the Presample, you must specify
the presample innovation or conditional variance variable names by using
the PresampleInnovationVariable or
PresampleVarianceVariable name-value
argument.
Since R2023a
Variable of Presample containing presample innovation paths εt, specified as one of the following data types:
String scalar or character vector containing a variable name in
Presample.Properties.VariableNamesVariable index (integer) to select from
Presample.Properties.VariableNamesA length
numprevarslogical vector, wherePresampleInnovationVariable(selects variablej) = truejPresample.Properties.VariableNames, andsum(PresampleInnovationVariable)is1
The selected variable must be a numeric matrix and cannot contain missing values (NaN).
If you specify presample innovation data by using the Presample name-value argument, you must specify PresampleInnovationVariable.
Example: PresampleInnovationVariable="StockRateInnov0"
Example: PresampleInnovationVariable=[false false true false] or PresampleInnovationVariable=3 selects the third table variable as the presample innovation variable.
Data Types: double | logical | char | cell | string
Since R2023a
Variable of Presample containing data for the presample conditional
variances
σt2,
specified as one of the following data types:
String scalar or character vector containing a variable name in
Presample.Properties.VariableNamesVariable index (positive integer) to select from
Presample.Properties.VariableNamesA logical vector, where
PresampleVarianceVariable(selects variablej) = truejPresample.Properties.VariableNames
The selected variable must be a numeric vector and cannot contain missing values
(NaNs).
If you specify presample conditional variance data by using the Presample name-value argument, you must specify PresampleVarianceVariable.
Example: PresampleVarianceVariable="StockRateVar0"
Example: PresampleVarianceVariable=[false false true false] or PresampleVarianceVariable=3 selects the third table variable as the presample conditional variance variable.
Data Types: double | logical | char | cell | string
Notes:
NaNvalues inY,E0, andV0indicate missing values.inferremoves missing values from specified data by list-wise deletion.For the presample,
inferhorizontally concatenatesE0andV0, and then it removes any row of the concatenated matrix containing at least oneNaN.For in-sample data
Y,inferremoves any row containing at least oneNaN.
This type of data reduction reduces the effective sample size and can create an irregular time series.
For numeric data inputs,
inferassumes that you synchronize the presample data such that the latest observations occur simultaneously.inferissues an error when any table or timetable input contains missing values.
Output Arguments
Conditional variances inferred from the response data
Y, returned as a numeric column vector or matrix.
infer returns V only
when you supply the input Y.
The dimensions of V and Y are
equivalent. If Y is a matrix, then the columns of
V are the inferred conditional variance paths
corresponding to the columns of Y.
Rows of V are periods corresponding to the periodicity
of Y.
Since R2023b
Inferred conditional variance
σt2
and innovation εt paths, returned
as a table or timetable, the same data type as Tbl1.
infer returns Tbl2 only
when you supply the input Tbl1. When
Mdl is an estimated model returned by
estimate, the returned, inferred innovations are
residuals.
Tbl2 contains the following variables:
The inferred conditional variance paths, which are in a
numobs-by-numpathsnumeric matrix, with rows representing observations and columns representing independent paths. Each path represents the continuation of the corresponding path of presample conditional variances inPresample.infernames the filtered conditional variance variable inTbl2responseName_VarianceresponseNameMdl.SeriesName. For example, ifMdl.SeriesNameisStockReturns,Tbl2contains a variable for the corresponding inferred conditional variance paths with the nameStockReturns_Variance.The inferred innovation paths, which are in a
numobs-by-numpathsnumeric matrix, with rows representing observations and columns representing independent paths. Each path corresponds to the input response path inTbl1and represents the continuation of the corresponding presample innovations path inPresample.infernames the inferred innovations variable inTbl2responseName_ResidualresponseNameMdl.SeriesName. For example, ifMdl.SeriesNameisStockReturns,Tbl2contains a variable for the corresponding inferred innovations paths with the nameStockReturns_Residual.All variables
Tbl1.
If Tbl1 is a timetable, row times of
Tbl1 and Tbl2 are
equal.
Algorithms
If you do not specify presample data (E0 and
V0, or Presample),
infer derives the necessary presample observations from
the unconditional, or long-run, variance of the offset-adjusted response process.
For all conditional variance model types, required presample conditional variances are the sample average of the squared disturbances of the offset-adjusted specified response data (
YorTbl1).For GARCH(P,Q) and GJR(P,Q) models, the required presample innovations are the square root of the average squared value of the offset-adjusted response data.
For EGARCH(P,Q) models, the required presample innovaitons are
0.
These specifications minimize initial transient effects.
References
[1] Bollerslev, T. “Generalized Autoregressive Conditional Heteroskedasticity.” Journal of Econometrics. Vol. 31, 1986, pp. 307–327.
[2] Bollerslev, T. “A Conditionally Heteroskedastic Time Series Model for Speculative Prices and Rates of Return.” The Review of Economics and Statistics. Vol. 69, 1987, pp. 542–547.
[3] Box, G. E. P., G. M. Jenkins, and G. C. Reinsel. Time Series Analysis: Forecasting and Control. 3rd ed. Englewood Cliffs, NJ: Prentice Hall, 1994.
[4] Enders, W. Applied Econometric Time Series. Hoboken, NJ: John Wiley & Sons, 1995.
[5] Engle, R. F. “Autoregressive Conditional Heteroskedasticity with Estimates of the Variance of United Kingdom Inflation.” Econometrica. Vol. 50, 1982, pp. 987–1007.
[6] Glosten, L. R., R. Jagannathan, and D. E. Runkle. “On the Relation between the Expected Value and the Volatility of the Nominal Excess Return on Stocks.” The Journal of Finance. Vol. 48, No. 5, 1993, pp. 1779–1801.
[7] Hamilton, J. D. Time Series Analysis. Princeton, NJ: Princeton University Press, 1994.
Version History
Introduced in R2012aIn addition to accepting input data (in-sample and presample) in numeric arrays,
infer accepts input data in tables or regular
timetables. When you supply data in a table or timetable, the following conditions
apply:
inferchooses the default in-sample response series on which to operate, but you can use the specified optional name-value argument to select a different series.If you specify optional presample innovation or conditional variance data to initialize the model, you must also specify the presample innovation or conditional variance series name.
inferreturns results in a table or timetable.
Name-value arguments to support tabular workflows include:
ResponseVariablespecifies the variable name of the response paths in the input data, from whichinferinfers conditional variances and innovations.Presamplespecifies the input table or timetable of presample innovation and conditional variance data.PresampleInnovationVariablespecifies the variable name of the innovation paths to select fromPresample.PresampleVarianceVariablespecifies the variable name of the conditional variance paths to select fromPresample.
See Also
Objects
Functions
MATLAB Command
You clicked a link that corresponds to this MATLAB command:
Run the command by entering it in the MATLAB Command Window. Web browsers do not support MATLAB commands.
Select a Web Site
Choose a web site to get translated content where available and see local events and offers. Based on your location, we recommend that you select: .
You can also select a web site from the following list
How to Get Best Site Performance
Select the China site (in Chinese or English) for best site performance. Other MathWorks country sites are not optimized for visits from your location.
Americas
- América Latina (Español)
- Canada (English)
- United States (English)
Europe
- Belgium (English)
- Denmark (English)
- Deutschland (Deutsch)
- España (Español)
- Finland (English)
- France (Français)
- Ireland (English)
- Italia (Italiano)
- Luxembourg (English)
- Netherlands (English)
- Norway (English)
- Österreich (Deutsch)
- Portugal (English)
- Sweden (English)
- Switzerland
- United Kingdom (English)