textureFeatures
Description
Examples
Load an X-ray image into the workspace as a medicalImage object. Visualize the image.
data = medicalImage("forearmXrayImage1.dcm");
I = data.Pixels;
figure
imshow(I,[])Draw two regions of interest (ROI) in the X-ray image. Create masks from the ROIs.
roi1 = drawassisted(Color="g"); roi2 = drawassisted(Color="r");
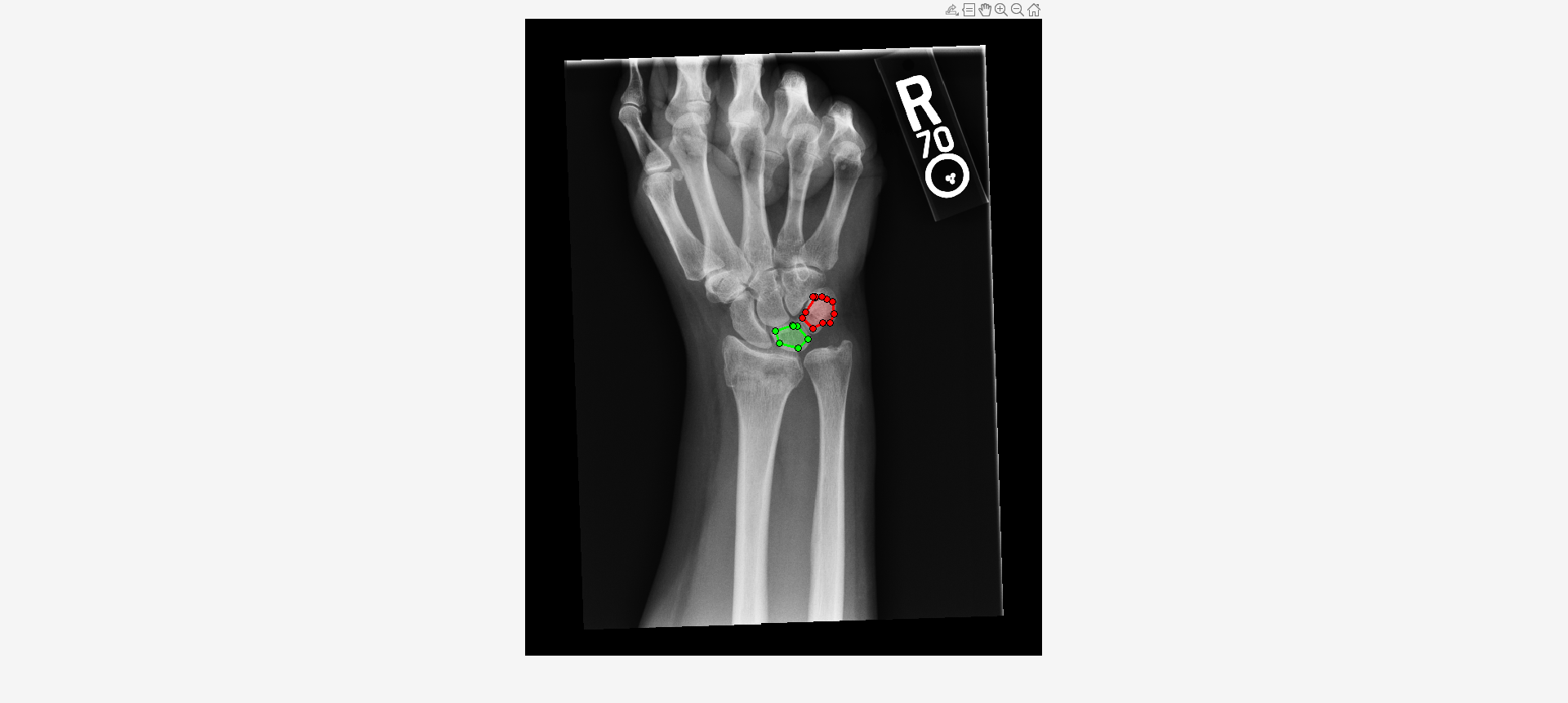
mask1 = createMask(roi1,I); mask2 = createMask(roi2,I);
Create an ROI label matrix, using different labels for the two ROIs. Create a medicalImage object of the ROI label data.
mask = zeros(size(I));
mask(mask1) = 1;
mask(mask2) = 2;
info = dicominfo("forearmXrayImage1.dcm");
roi = medicalImage(mask,info);Create a radiomics object from the X-ray image data and ROI label data.
R = radiomics(data,roi)
R =
radiomics with properties:
Data: [1×1 medicalImage]
ROILabel: [1×1 medicalImage]
Resample: 0
Resegment: 1
Discretize: 1
DiscretizeIVH: 1
ResampledVoxelSpacing: []
DataResampleMethod: []
MaskResampleMethod: []
ResegmentationRange: []
ExcludeOutliers: 1
DiscreteBinSizeOrBinNumber: []
DiscreteMethod: 'FixedBinNumber'
DiscreteIVHBinSizeOrBinNumber: []
DiscreteIVHMethod: 'FixedBinNumber'
Compute texture features for both ROIs.
T = textureFeatures(R)
T=2×137 table
"1" 0.0025 57.1732 675.8153 11.0685 8.6729 46.5654 4.5233 114.3463 2.5804e+03 7.4309 5.9926e-04 122.8478 8.6729 0.1971 0.9492 0.1125 0.9951 0.1163 0.9090 3.8832e+03 2.5804e+03 1.3078e+05 23618430 -0.2975 0.9893 0.0016 57.1739 675.8519 11.5961
"2" 0.0021 95.4544 1.0353e+03 11.4294 7.9161 41.0320 4.4079 190.9088 4.0368e+03 7.7995 4.6008e-04 104.5714 7.9161 0.2109 0.9534 0.1239 0.9958 0.1264 0.9495 1.0095e+04 4.0368e+03 -1.9066e+05 48382976 -0.3299 0.9944 0.0016 95.4542 1.0354e+03 11.8986
Import a computed tomography (CT) image volume and the corresponding ROI mask volume from the IBSI validation data set [1][2][3] as medicalVolume objects.
unzip("CTImageMaskNIfTI.zip") data = medicalVolume("CT_image.nii.gz"); roi = medicalVolume("CT_mask.nii.gz");
Visualize a slice of the CT image volume and the corresponding ROI.
figure
imshowpair(data.Voxels(:,:,20),roi.Voxels(:,:,20),"montage")
Create a radiomics object, using the CT image volume and ROI mask volume, with default preprocessing options.
R = radiomics(data,roi)
R =
radiomics with properties:
Data: [1×1 medicalVolume]
ROILabel: [1×1 medicalVolume]
Resample: 1
Resegment: 1
Discretize: 1
DiscretizeIVH: 1
ResampledVoxelSpacing: 1
DataResampleMethod: 'linear'
MaskResampleMethod: 'linear'
ResegmentationRange: []
ExcludeOutliers: 1
DiscreteBinSizeOrBinNumber: []
DiscreteMethod: 'FixedBinNumber'
DiscreteIVHBinSizeOrBinNumber: []
DiscreteIVHMethod: 'FixedBinNumber'
Compute the grey level co-occurrence matrix (GLCM) texture features of the ROI in the 2-D resampled CT image volume.
I = textureFeatures(R,Type=["GLCM","GLRLM"],SubType="2D")
I=1×83 table
LabelID JointMaximumAveraged2D JointAverageAveraged2D JointVarianceAveraged2D JointEntropyAveraged2D DifferenceAverageAveraged2D DifferenceVarianceAveraged2D DifferenceEntropyAveraged2D SumAverageAveraged2D SumVarianceAveraged2D SumEntropyAveraged2D AngularSecondMomentAveraged2D ContrastAveraged2D DissimilarityAveraged2D InverseDifferenceAveraged2D NormalisedInverseDifferenceAveraged2D InverseDifferenceMomentAveraged2D NormalisedInverseDifferenceMomentAveraged2D InverseVarianceAveraged2D CorrelationAveraged2D AutoCorrelationAveraged2D ClusterTendencyAveraged2D ClusterShadeAveraged2D ClusterProminenceAveraged2D InformationCorrelation1Averaged2D InformationCorrelation2Averaged2D JointMaximumSliceMerged2D JointAverageSliceMerged2D JointVarianceSliceMerged2D JointEntropySliceMerged2D DifferenceAverageSliceMerged2D DifferenceVarianceSliceMerged2D DifferenceEntropySliceMerged2D SumAverageSliceMerged2D SumVarianceSliceMerged2D SumEntropySliceMerged2D AngularSecondMomentSliceMerged2D ContrastSliceMerged2D DissimilaritySliceMerged2D InverseDifferenceSliceMerged2D NormalisedInverseDifferenceSliceMerged2D InverseDifferenceMomentSliceMerged2D NormalisedInverseDifferenceMomentSliceMerged2D InverseVarianceSliceMerged2D CorrelationSliceMerged2D AutoCorrelationSliceMerged2D ClusterTendencySliceMerged2D ClusterShadeSliceMerged2D ClusterProminenceSliceMerged2D InformationCorrelation1SliceMerged2D InformationCorrelation2SliceMerged2D ShortRunsEmphasisAveraged2D LongRunsEmphasisAveraged2D LowGrayLevelRunEmphasisAveraged2D HighGrayLevelRunEmphasisAveraged2D ShortRunLowGrayLevelEmphasisAveraged2D ShortRunHighGrayLevelEmphasisAveraged2D LongRunLowGrayLevelEmphasisAveraged2D LongRunHighGrayLevelEmphasisAveraged2D GrayLevelNonUniformityAveraged2D NormalisedGrayLevelNonUniformityAveraged2D RunLengthNonUniformityAveraged2D NormalisedRunLengthNonUniformityAveraged2D RunPercentageAveraged2D GrayLevelVarianceAveraged2D RunLengthVarianceAveraged2D RunEntropyAveraged2D ShortRunsEmphasisSliceMerged2D LongRunsEmphasisSliceMerged2D LowGrayLevelRunEmphasisSliceMerged2D HighGrayLevelRunEmphasisSliceMerged2D ShortRunLowGrayLevelEmphasisSliceMerged2D ShortRunHighGrayLevelEmphasisSliceMerged2D LongRunLowGrayLevelEmphasisSliceMerged2D LongRunHighGrayLevelEmphasisSliceMerged2D GrayLevelNonUniformitySliceMerged2D NormalisedGrayLevelNonUniformitySliceMerged2D RunLengthNonUniformitySliceMerged2D NormalisedRunLengthNonUniformitySliceMerged2D RunPercentageSliceMerged2D GrayLevelVarianceSliceMerged2D RunLengthVarianceSliceMerged2D RunEntropySliceMerged2D
_______ ______________________ ______________________ _______________________ ______________________ ___________________________ ____________________________ ___________________________ ____________________ _____________________ ____________________ _____________________________ __________________ _______________________ ___________________________ _____________________________________ _________________________________ ___________________________________________ _________________________ _____________________ _________________________ _________________________ ______________________ ___________________________ _________________________________ _________________________________ _________________________ _________________________ __________________________ _________________________ ______________________________ _______________________________ ______________________________ _______________________ ________________________ _______________________ ________________________________ _____________________ __________________________ ______________________________ ________________________________________ ____________________________________ ______________________________________________ ____________________________ ________________________ ____________________________ ____________________________ _________________________ ______________________________ ____________________________________ ____________________________________ ___________________________ __________________________ _________________________________ __________________________________ ______________________________________ _______________________________________ _____________________________________ ______________________________________ ________________________________ __________________________________________ ________________________________ __________________________________________ _______________________ ___________________________ ___________________________ ____________________ ______________________________ _____________________________ ____________________________________ _____________________________________ _________________________________________ __________________________________________ ________________________________________ _________________________________________ ___________________________________ _____________________________________________ ___________________________________ _____________________________________________ __________________________ ______________________________ ______________________________ _______________________
"1" 0.025292 10.263 9.9482 6.9415 2.6834 4.353 2.8719 20.525 27.92 4.3458 0.011452 11.872 2.6834 0.39071 0.8891 0.3042 0.97287 0.30871 0.39971 111.04 27.92 -17.682 2401.8 -0.10434 0.66674 0.021298 10.262 9.9491 7.1238 2.678 4.5887 2.9277 20.525 27.962 4.3981 0.0098741 11.835 2.678 0.39129 0.8893 0.3049 0.97295 0.30908 0.40201 111.06 27.962 -18.094 2408.5 -0.052692 0.55415 0.91083 1.4376 0.023078 119.32 0.021376 108.52 0.030767 173.59 75.51 0.087674 710.23 0.79414 0.896 12.076 0.15199 4.243 0.91189 1.4316 0.023109 119.34 0.021486 108.64 0.030465 172.87 301.76 0.087414 2836.1 0.79414 0.896 12.083 0.15192 4.2827
[1] Vallières, Martin, Carolyn R. Freeman, Sonia R. Skamene, and Issam El Naqa. “A Radiomics Model from Joint FDG-PET and MRI Texture Features for the Prediction of Lung Metastases in Soft-Tissue Sarcomas of the Extremities.” The Cancer Imaging Archive, 2015. https://doi.org/10.7937/K9/TCIA.2015.7GO2GSKS.
[2] Vallières, M, C R Freeman, S R Skamene, and I El Naqa. “A Radiomics Model from Joint FDG-PET and MRI Texture Features for the Prediction of Lung Metastases in Soft-Tissue Sarcomas of the Extremities.” Physics in Medicine and Biology 60, no. 14 (July 7, 2015): 5471–96. https://doi.org/10.1088/0031-9155/60/14/5471.
[3] Clark, Kenneth, Bruce Vendt, Kirk Smith, John Freymann, Justin Kirby, Paul Koppel, Stephen Moore, et al. “The Cancer Imaging Archive (TCIA): Maintaining and Operating a Public Information Repository.” Journal of Digital Imaging 26, no. 6 (December 2013): 1045–57. https://doi.org/10.1007/s10278-013-9622-7.
Input Arguments
Name-Value Arguments
Specify optional pairs of arguments as
Name1=Value1,...,NameN=ValueN, where Name is
the argument name and Value is the corresponding value.
Name-value arguments must appear after other arguments, but the order of the
pairs does not matter.
Example: textureFeatures(R,Type="GLCM",SubType="2D") computes the GLCM
texture features as the average over 2-D slices in the 2-D resampled data.
Category of texture features to compute, specified as one or more of these options.
"GLCM"— Grey level co-occurrence matrix"GLRLM"— Grey level run length matrix"GLSZM"— Grey level size zone matrix"GLDZM"— Grey level distance zone matrix"NGTDM"— Neighbourhood grey tone difference matrix"NGLDM"— Neighbourhood grey level dependence matrix"all"
If you specify "all", the function computes every
category of texture features. For more information on which specific texture features
each category includes, see IBSI Standard and Radiomics Function Feature Correspondences.
Data Types: char | string
Resampling from which to compute texture features, specified as one of these options.
"2D"— Computes features for each 2-D slice in the 2-D resampled data and averages them. This is the default option for 2-D data, when theDataandROILabelproperties of theradiomicsobjectRare 2-D matrices ormedicalImageobjects."2.5D"— Computes features after merging all 2-D slices in the 2-D resampled data."3D"— Computes features for the entire 3-D volume in the 3-D resampled data. This is the default option for 3-D data, when theDataandROILabelproperties of theradiomicsobjectRare 3-D arrays ormedicalVolumeobjects. 3-D resampling is not applicable for 2-D data."all"— Computes features for all applicable options.
For 3-D data, when you perform 3-D resampling of the volume, the function makes the voxel spacing along all three spatial dimensions isotropic. However, when you perform 2-D resampling of the volume, the function makes the voxel spacing along only the x- and y-dimensions isotropic, while retaining the voxel spacing of the input volume along the z-dimension.
Data Types: char | string
Output Arguments
Texture features, returned as a table. The first column in T is
LabelID. The subsequent columns are the texture features. Each row
of the table corresponds to an ROI. For more details on which texture features are
computed in each Type and SubType, see IBSI Standard and Radiomics Function Feature Correspondences.
Version History
Introduced in R2023bCompute texture features for multiple regions of interest (ROIs) in a medical image or medical volume.
Compute one or more types of texture features by specifying one or more options in the
name-value argument Type.
MATLAB Command
You clicked a link that corresponds to this MATLAB command:
Run the command by entering it in the MATLAB Command Window. Web browsers do not support MATLAB commands.
Select a Web Site
Choose a web site to get translated content where available and see local events and offers. Based on your location, we recommend that you select: .
You can also select a web site from the following list
How to Get Best Site Performance
Select the China site (in Chinese or English) for best site performance. Other MathWorks country sites are not optimized for visits from your location.
Americas
- América Latina (Español)
- Canada (English)
- United States (English)
Europe
- Belgium (English)
- Denmark (English)
- Deutschland (Deutsch)
- España (Español)
- Finland (English)
- France (Français)
- Ireland (English)
- Italia (Italiano)
- Luxembourg (English)
- Netherlands (English)
- Norway (English)
- Österreich (Deutsch)
- Portugal (English)
- Sweden (English)
- Switzerland
- United Kingdom (English)